Developing Multi-Step Workflows
Overview
Teaching: 0 min
Exercises: 0 minQuestions
How can we expand to a multi-step workflow?
Iterative workflow development
Workflows as dependency graphs
How to use sketches for workflow design?
Objectives
explain that a workflow is a dependency graph
use cwlviewer online
generate Graphviz diagram using cwltool
exercise with the printout of a simple workflow; draw arrows on code; hand draw a graph on another sheet of paper
recognise that workflow development can be iterative i.e. that it doesn’t have to happen all at once
understand the flow of data between tools
Multi-Step Workflow
In the previous episode a single step workflow was shown. To make a multi-step workflow, you add more entries to the steps field.
In this episode, the workflow is extended with the next two steps of the RNA-sequencing analysis.
The next two steps are triming the reads and alignment of the trimmed reads.
We will be using the cutadapt and STAR tools for these tasks.
rna_seq_workflow.cwl
cwlVersion: v1.2
class: Workflow
inputs:
rna_reads_forward:
type: File
format: https://edamontology.org/format_1930 # FASTQ
rna_reads_reverse:
type: File
format: https://edamontology.org/format_1930 # FASTQ
ref_genome: Directory
gene_model: File
steps:
quality_control_forward:
run: bio-cwl-tools/fastqc/fastqc_2.cwl
in:
reads_file: rna_reads_forward
out: [html_file]
quality_control_reverse:
run: bio-cwl-tools/fastqc/fastqc_2.cwl
in:
reads_file: rna_reads_reverse
out: [html_file]
trim_low_quality_bases:
run: bio-cwl-tools/cutadapt/cutadapt-paired.cwl
in:
reads_1: rna_reads_forward
reads_2: rna_reads_reverse
minimum_length: { default: 20 }
quality_cutoff: { default: 20 }
out: [ trimmed_reads_1, trimmed_reads_2, report ]
mapping_reads:
requirements:
ResourceRequirement:
ramMin: 9000
run: bio-cwl-tools/STAR/STAR-Align.cwl
in:
RunThreadN: {default: 4}
GenomeDir: ref_genome
ForwardReads: trim_low_quality_bases/trimmed_reads_1
ReverseReads: trim_low_quality_bases/trimmed_reads_2
OutSAMtype: {default: BAM}
SortedByCoordinate: {default: true}
OutSAMunmapped: {default: Within}
Overhang: { default: 36 } # the length of the reads - 1
Gtf: gene_model
out: [alignment]
outputs:
alignments:
type:
- File
- File[]
outputSource: mapping_reads/alignment
The workflow file shows the first 3 steps of the RNA-seq analysis: quality_control, mapping_reads and index_alignment.
The index_alignment step uses the alignment output of the mapping_reads step.
You do this by referencing the output of the mapping_reads step in the in field of the index_alignment step.
This is similar to referencing the outputs of the different steps in the outputs section.
The mapping_reads step needs some extra information beyond the inputs from the other steps, which is done by providing default values. If you want, you can read the bio-cwl-tools/STAR/STAR-Align.cwl file to see how these extra inputs are transformed into command line options to the STAR program.
This information is provided in the in field.
To run the tool better, it needs more RAM than the default. So there is a requirements entry inside the mapping_reads step definition with a ResourceRequirement to allocate a minimum of 9000 MiB of RAM.
The newly added mapping_reads step also need an input not provided by any of our other steps, therefore an additional workflow-level input is added: a directory that contains the reference genome necessary for the mapping.
This ref_genome is added in the inputs field of the workflow and in the YAML input file, workflow_input.yml.
workflow_input.yml
rna_reads_forward:
class: File
location: rnaseq/GSM461177_1_subsampled.fastqsanger
format: https://edamontology.org/format_1930 # FASTQ
rna_reads_reverse:
class: File
location: rnaseq/GSM461177_2_subsampled.fastqsanger
format: https://edamontology.org/format_1930 # FASTQ
ref_genome:
class: Directory
location: rnaseq/dm6-STAR-index
gene_model:
class: File
location: rnaseq/Drosophila_melanogaster.BDGP6.87.gtf
Exercise
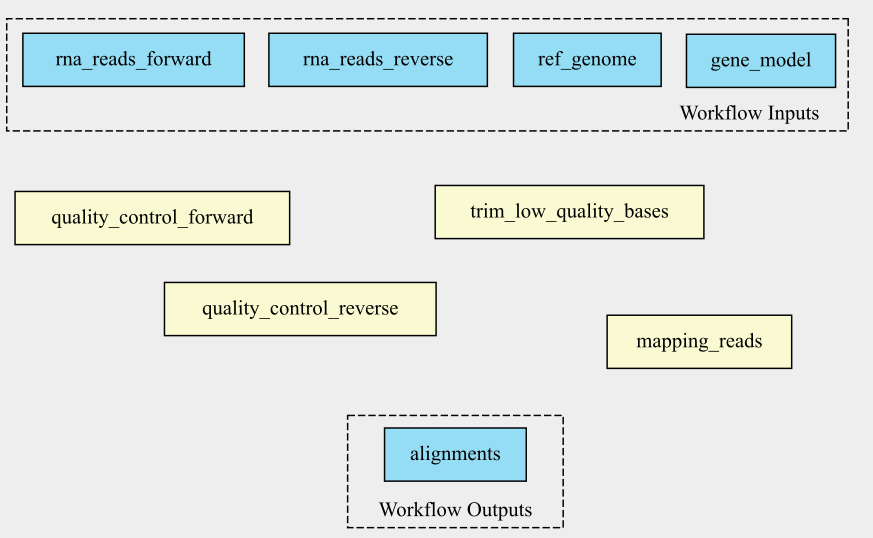
Draw the connecting arrows in the following graph of our workflow. Also, provide the outputs/inputs of the different steps. You can use for example Paint or print out the graph.
Solution
To find out how the inputs and the steps are connected to each other, you look at the
infield of the different steps.
Iterative working
Working on a workflow is often not something that happens all at once. Sometimes you already have a shell script ready that can be converted to a CWL workflow. Other times it is similar to this tutorial, you start with a single-step workflow and extend it to a multi-step workflow. This is all iterative working, a continuous work in progress.
Visualising a workflow
A CWL workflow is a directed acyclic graph (DAG). This means that:
- The workflow has a certain direction, from workflow inputs to step inputs, from step outputs to other step inputs, and from step outputs to workflow outputs and
- The workflow definition has no cycles. A CWL workflow is a dependency graph. Each input for a step in the workflow depends on either a workflow-level input or a particular output from another step.
To visualise a workflow, a graph can be used. This can be done before a CWL script is written to visualise how the different steps connect to each other.
It is also possible to make a graph after the CWL script has been written. This graph can be generated using online tools or the build-in function in cwltool.
When a graph is generated, it can be used to visualise the steps taken and could make it easier to explain a workflow to other researchers.
From CWL script to graph
In this example the workflow is already made, so the graph can be generated using cwlviewer online or using cwltool.
First, let’s have a look at cwlviewer. To use this tool, the workflow has to be put in a GitHub, GitLab or Git repository.
To view the graph of the workflow enter the URL and click Parse Workflow. The cwlviewer displays the workflow as a graph, starting with the input.
Then the different steps are shown, each with their input(s) and output(s). The steps are linked to eachother using arrows accompanied by the input of the next step.
The graph ends with the workflow outputs.
The graph of the RNA-seq workflow looks a follows:
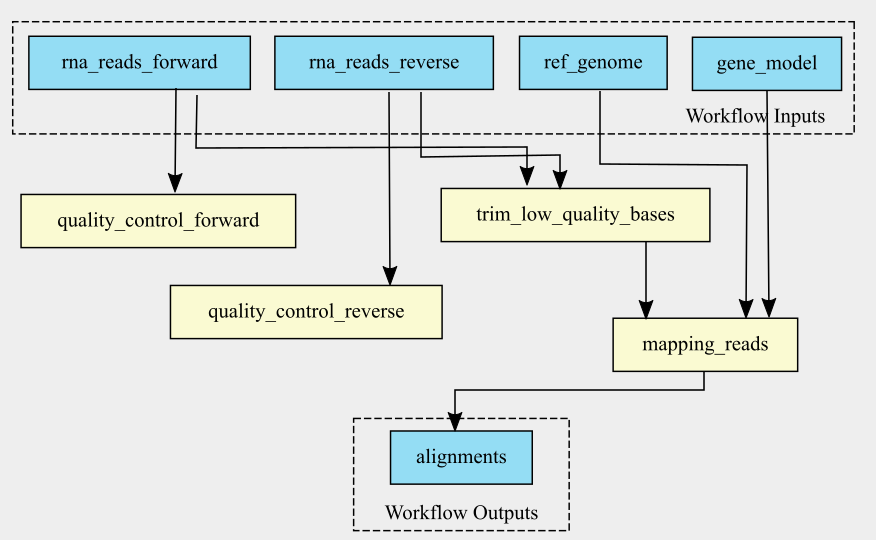
It is also possible to generate the graph in the command line. cwltool has a function that makes a graph.
The --print-dot option will print a file suitable for Graphviz dot program. This is the command to generate a Scalable Vector Graphic (SVG) file:
cwltool --print-dot rna_seq_workflow.cwl | dot -Tsvg > workflow_graph.svg
The resulting SVG file displays the same graph as the one in the cwlviewer. The SVG file can be opened in any web browser and in Inkscape, for example.
Visualisation in VSCode
Benten is an extension in Visual Studio Code (VSCode) that among other things visualises a workflow in a graph. When Benten is installed in VSCode, the tool can be used to visualise the workflow. In the top-right corner of the VSCode window the CWL viewer can be opened, see the screenshot below.
.png)
In VSCode/Benten the inputs are shown in green, the steps in blue and the outputs in yellow. This graph looks a little bit different from the graph made with cwlviewer or cwltool.
The graph by VSCode/Benten doesn’t show the output-input names between the different steps.
.png)
Key Points
A multi-step workflow has multiple entries under the
stepssectionWorkflow development can be an iterative process
A CWL workflow can be represented as a dependency graph, either to explain your workflow or as a planning tool